The Facility provides high-quality short-read library preparations and sequencing with the Illumina NovaSeq 6000 and MiSeq sequencers. We also provide high-fidelity (HiFi) long-read DNA and RNA sequencing with the PacBio Revio sequencer, and we operate an Oxford Nanopore PromethION 24 as an open access platform and to offer long-reads library preparations and sequencing services.
Short Read Sequencing
Technology
Illumina sequencing produces data at rates orders of magnitude faster than Sanger sequencing, but generates shorter sequences (50 to 300 bases each). Millions to billions of DNA molecules are covalently attached to the surface of a treated glass flowcell, and each individual molecule is amplified in situ to generate a “cluster” that contains about 1000 copies. Sequences are read from newly synthesised DNA copied from these targets by incorporation of fluorescently-tagged nucleotide analogues, illumination with a laser and optical capture of signal from all the amplified clusters simultaneously. Edinburgh Genomics offers the low (MiSeq) and very high (NovaSeq 6000) technologies.
The NovaSeq is capable of generating many tens of gigabases per day. These data can be:
- assembled de novo to predict the sequence of a new genome or transcriptome
- aligned to a reference genome to identify single nucleotide or insertion-deletion polymorphisms
- used to count the abundance of species in a DNA mix. The mixes can be the 3' ends of transcripts, small RNAs such as microRNAs or DNA fragments generated by chromatin immunoprecipitation (ChIP) experiments
We can multiplex many samples per run, so whether you want a “small” number of short reads or hundreds of gigabases, we can deliver.

Applications
Our sequencing instrumentation is very versatile: if the sample is nucleic acid, we can transform it to a sequencable library. Data generated by Edinburgh Genomics have been applied to questions in a wide range of fields:
| Genome assembly | SNP discovery | CNV mapping | Genome rearrangements | Genome architecture | Epigenetic modification | Genetic mapping | Quantitative trait genetics | Population genetics | Metagenomics | Metagenetics | Mutation screening | Synthetic biology | Transcriptome assembly | Gene expression | Transcriptional regulation | Phenotyping of mutants | Pathogen discovery | Evolutionary genomics | Host-pathogen interaction | Ecotoxicogenomics |
The Illumina, PacBio, and Oxford Nanopore instruments are suited to a wide range of approaches, including:
DNA-Seq:
- De novo genome sequencing
- Genome resequencing
- Targeted resequencing
- Exome sequencing
- Amplicon sequencing
- ChIP sequencing (and variants, including 5C)
- Methylome sequencing
RNA-Seq:
- Stranded RNA sequencing
- Single Cell RNA sequencing
- Small RNA sequencing
- Total RNA sequencing

Long Read Sequencing
Platforms
The PacBio Revio and Oxford Nanopore PromethION are capable of generating many tens of gigabases per day. These data can be:
- assembled de novo to predict the sequence of a new genome or full length transcriptome
- aligned to a reference genome to identify single nucleotide, insertion-deletion polymorphisms, and structural variants
- the PromethION can generate RNA sequence directly
We can multiplex many samples per run, making the technologies very flexible.
We work closely with PacBio and Oxford Nanopore to offer the best long read technologies for your research.


Technology
PacBio Revio
The PacBio Revio is based on PacBio’s single molecule, real-time technology (SMRT) and delivers the highly accurate long HiFi reads. This platform delivers long reads, high consensus accuracy with HiFi and epigenetic characterisation and is ideal for generating high-quality whole genome de novo assemblies and RNA isoform characterization.
Contact us to take advantage of the latest PacBio long-read technology for applications including de novo genome assemblies, structural variation, ISO-Seq, long amplicons, single cell transcriptomics, full 16S rRNA and methylome detection.
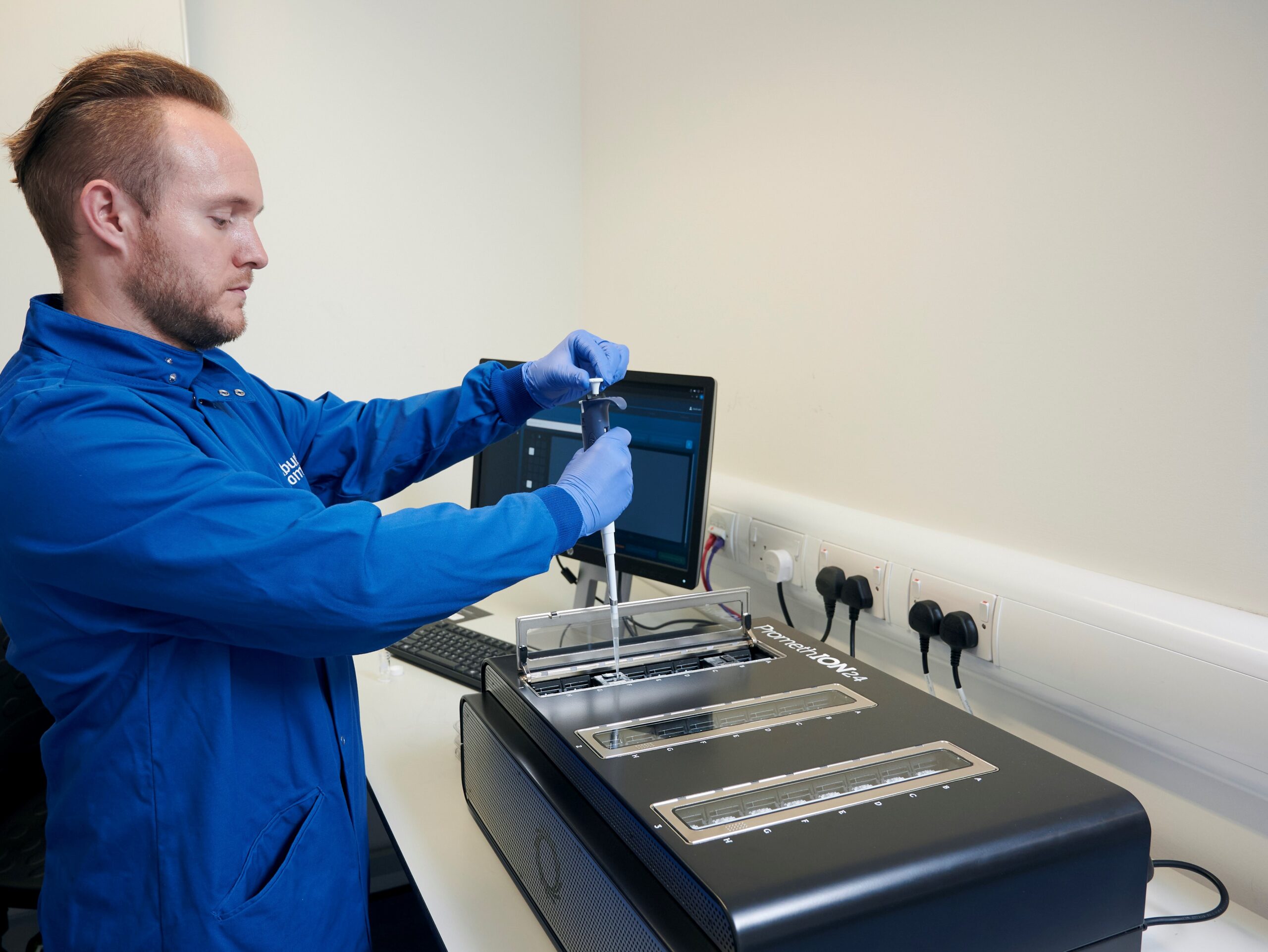
Oxford Nanopore PromethION
The ONT PromethION offers real-time, long-read, direct DNA and RNA sequencing technology at a large scale. It offers the same pore-based sequencing as the MinION, but with many more pores for a much higher throughput and lower cost per base.



